How to set up the sample sheet for a PhiX validation run on the MiSeq using IEM - Illumina Knowledge

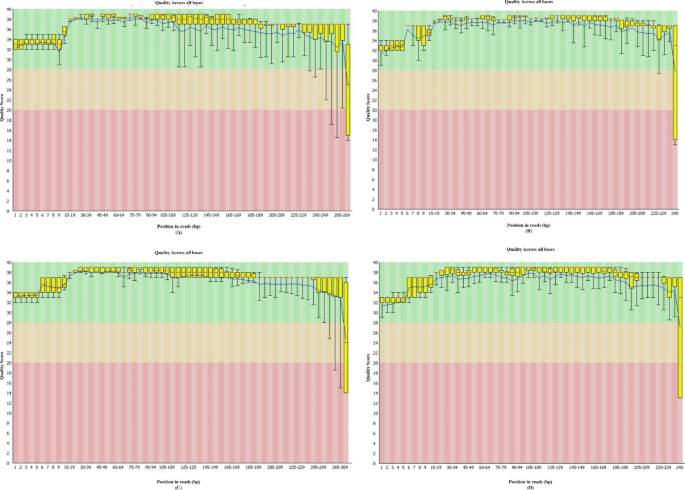
I had a Metagenomic 16s rRNA run on MiSeq v3 kit 2 x300 cycles. with good PF >89% but low quality q30 <40%. Kindly advice what could get wrong? | ResearchGate

How to set up the sample sheet for a PhiX validation run on the MiSeq using IEM - Illumina Knowledge
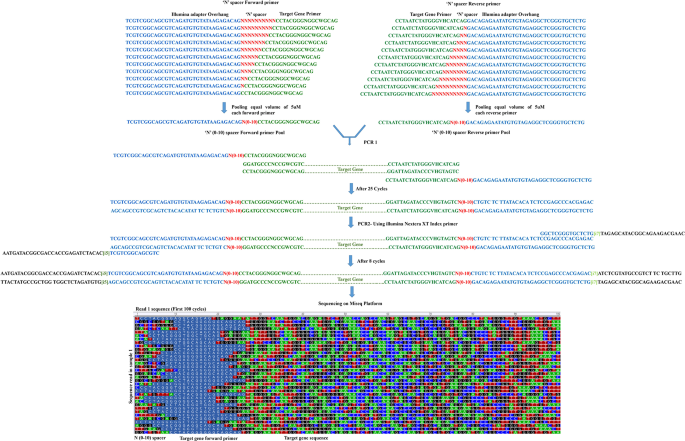
High-quality single amplicon sequencing method for illumina MiSeq platform using pool of 'N' (0–10) spacer-linked target specific primers without PhiX spike-in | BMC Genomics | Full Text